Abstract:
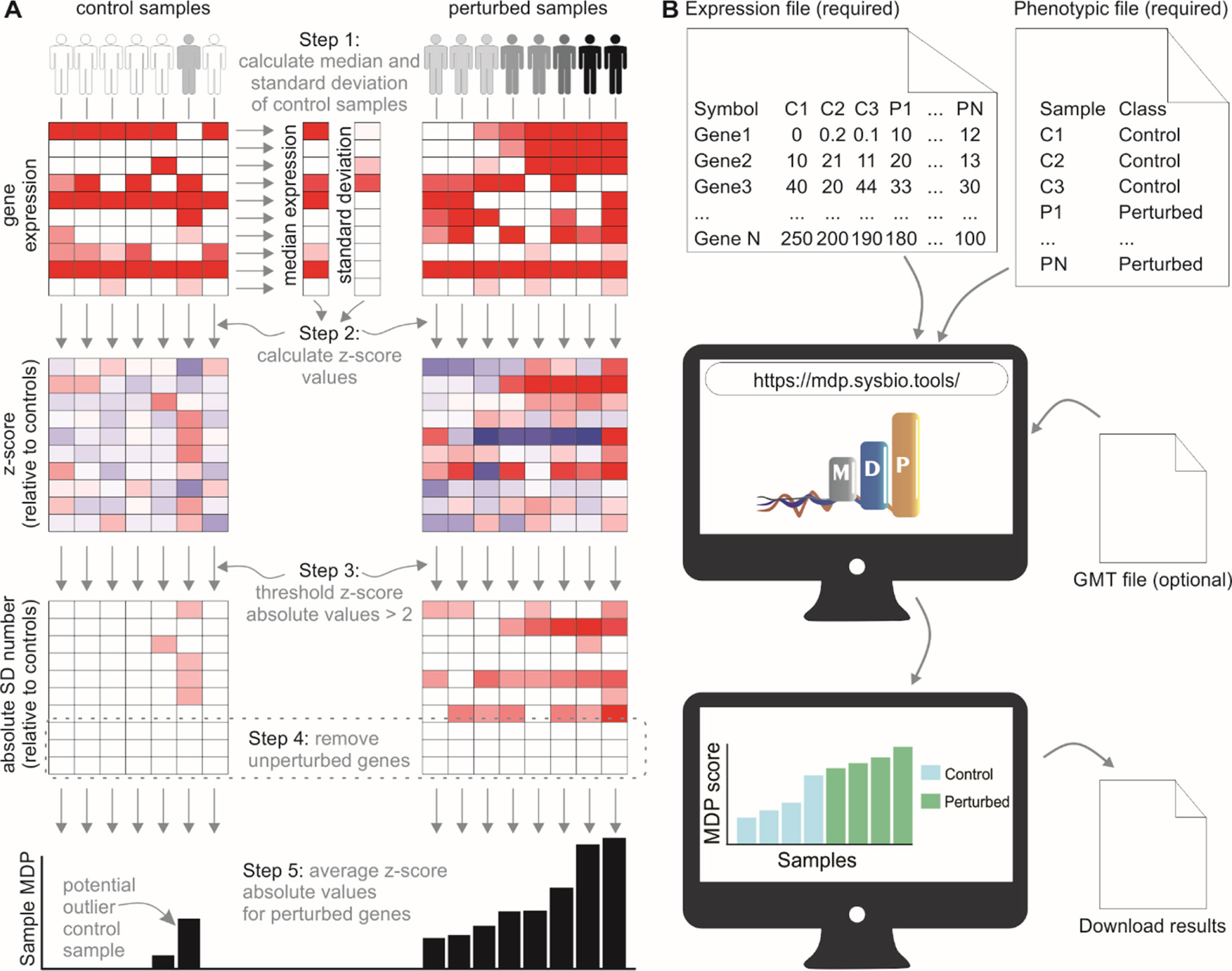
Transcriptome analyses have increased our understanding of the molecular mechanisms underlying human diseases. Most approaches aim to identify significant genes by comparing their expression values between healthy subjects and a group of patients with a certain disease. Given that studies normally contain few samples, the heterogeneity among individuals caused by environmental factors or undetected illnesses can impact gene expression analyses. We present a systematic analysis of sample heterogeneity in a variety of gene expression studies relating to inflammatory and infectious diseases and show that novel immunological insights may arise once heterogeneity is addressed. The perturbation score of samples is quantified using nonperturbed subjects (i.e., healthy subjects) as a reference group. Such a score allows us to detect outlying samples and subgroups of diseased patients and even assess the molecular perturbation of single cells infected with viruses. We also show how removal of outlying samples can improve the “signal” of the disease and impact detection of differentially expressed genes. The method is made available via the mdp Bioconductor R package and as a user-friendly webtool, webMDP, available at http://mdp.sysbio.tools.
Figure: